-it was discovered in a study of blood clotting factors in 1968
-it is used by food chefs as a bonding agent for proteins
Ajinomoto Food Ingredients
Since this enzyme is a common one, it is isolated using a fermentation process, and you basically sprinkle it on different meats that you want to bond together. It's really popular in the "molecular gastronomy" field - the people that make nitrogen-frozen strawberries for desserts and use blow torches a lot.
-In patients with celiac, gluten comes into contact with transglutaminase and the enzyme modifies the gluten. This reaction causes an elevated immune response that is poorly understood, but the results are the synthesis of far too many antibodies that tell the body transglutaminase is toxic and must be destroyed. Thus, in celiac patients, they are lacking the ability to digest gluten, and the immune response causes inflammation in the digestive system which also hampers digestion as a whole, glutenacious or not.
-use as a surgical glue and tumor treatment is being investigated
So what does this have to do with malaria, you ask? Well...even I really don't know yet. But the really interesting part of this enzyme is that it has a drastic conformational change that occurs with the addition of calcium and a given substrate (think gluten as as example)
 So...the best way that I found to think about these two enzymes is using the "Heads, Shoulders Knees and Toes" game. Red is the head, orange is the shoulders, green is the knees, blue is the toes. Thus, when the enzyme is in "closed" position, the head is really close to the toes, but in open conformation, they are really far away. This is about as dramatic a protein change can get.
So...the best way that I found to think about these two enzymes is using the "Heads, Shoulders Knees and Toes" game. Red is the head, orange is the shoulders, green is the knees, blue is the toes. Thus, when the enzyme is in "closed" position, the head is really close to the toes, but in open conformation, they are really far away. This is about as dramatic a protein change can get.What is even more important is how close the two ends of the protein are to one another. Many proteins are made of many different parts that are bonded together, but this one is basically one long string of amino acids, which in the above picture is represented by the ribbon and stringy things (fittingly, this is called a ribbon model).
The picture below is more 3D - you can see a bit more of how the molecules fit into space. You can also download a special program that will let you rotate this structure around in 3D space to try and better understand how it works.
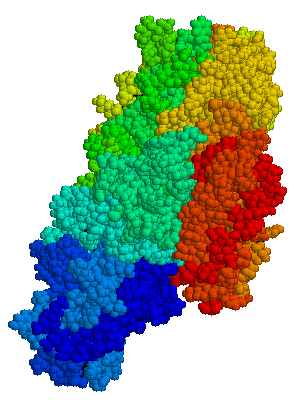
However, with this picture, you can't where the protein begins and ends...which is why the lab is excited about this enzyme. If you look closely at the ribbon model, you can see that one end of the protein is actually really close to the other end of the protein when the enzyme is in the "closed" position and really far apart when they are in the open position.
There are some really interesting things we could do with this in terms of making it into a molecular tool, but first I have to do some background research into how the folding and unfolding actually works, as well as how to detect how it is working during an experiment, and if that can be done, see if I can control the movement with some small molecule...as in, instead of the enzyme going back and forth between positions due to enzymatic activity, I would add a different non-related substrate and get it to do the same thing.
First, however, I have to map out some experiments to get things started.
1 comment:
You need to learn how to get to the point, and make things easily understood, this just seems like a jumbled mess of nothing to me....
Post a Comment